Hello,
I’ve just noticed a curious behaviuor and after thinking about it I believe it is intended, but I wanted to verify it and ask how to deal with it.
I had a subset of unstructured mesh resampled to image for an external analysis. In serial the procedure worked fine, but when run in parallel I noticed there are additional points and replotting the file in an external toolkit was showing an offset.
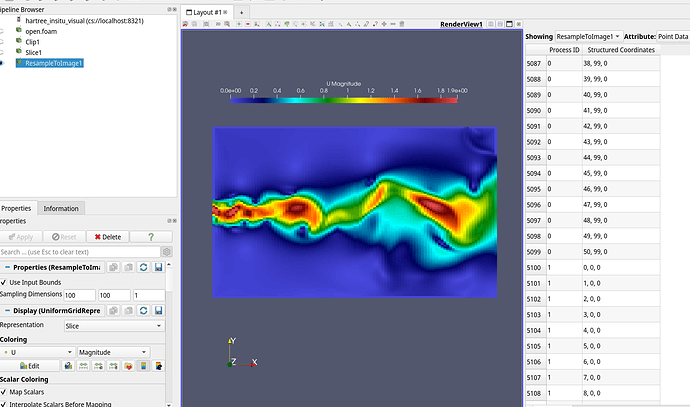
After investigating it more closely, I realised that the points originating from Resample to Image filter are broken up into different processors and the additional points are presumably the repetitions so e.g for 2 processes and 100x100 image the structured coordinate 50 is 0 in the next processor.
But if I use Save Data on the filter I lose Process ID and structured coordinate. My questions:
- Is it possible to recover the serial behaviour whilst working in parallel e.g. 100x100x1 should return 10k points rather than 11k?
- If not, how to reconstruct the image correctly when using Save Data?
I am ob obviously doing this for a large number of time step so need to automate. Would appreciate suggestions. Thanks.