Hi,
I am currently using the python Paraview API to analyze CFD results (.vti files), where I just want to do simple operations and visualizations of the cell data (velocities, pressure etc.). To do this, I am using the XMLImageDataReader to read the .vti files (each time step has its own .vti file) and servermanager.Fetch() to get the data loaded into memory for processing and extracting the variables of interest.
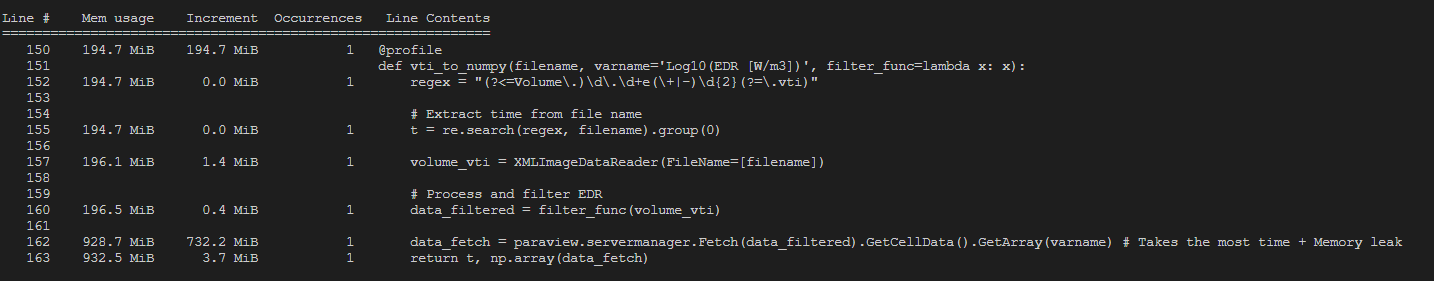
However, it seems that, once I extracted the variable of interest from the fetched data (using GetCellData() and GetArray()) the memory used by servermanager.Fetch() is not released. For each new file that is fetched, the memory usage just increases with that file size (approx 500MB per file). Even though I am using Paraviews Calculator and Threshold filters to extract only the cell values that actually contain a fluid (filter function passed directly to Fetch), it seems that still the complete file is loaded and never released from memory. Since the final size of my numpy array, which contains the values of interest, is only a ~3 MB, I expected the memory would be released after the iteration to the next file. However, it seems all files are kept in memory and are only released when the program terminates. For large data sets, this causes Python to crash due to a memory error.
To solve this problem I tried:
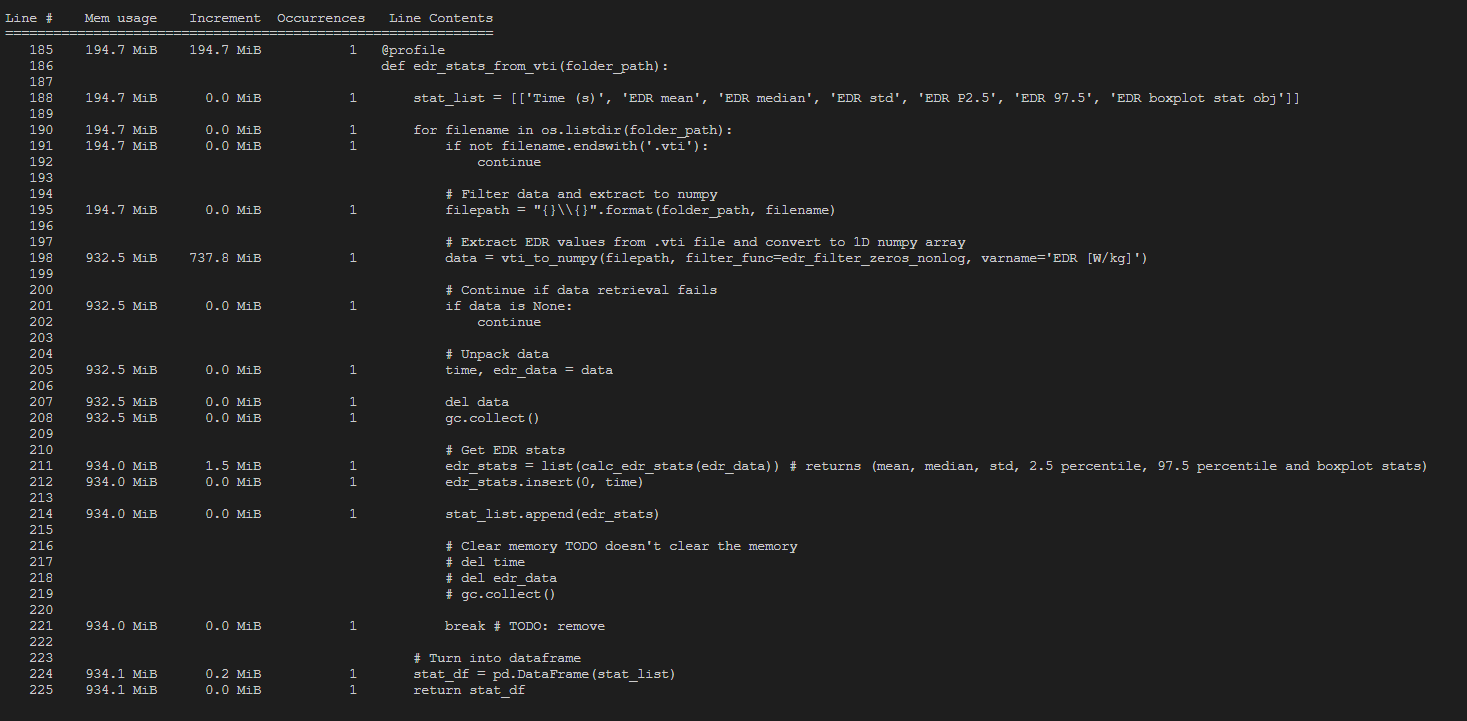
- to use Python’s Del operator and the garbage collector modules to see if I can clear the memory, but that didn’t work
- to modify the filter functions passed to Fetch to only fetch as little data as possible, but still the complete .vti file is loaded in memory and not released.
- upgrading to Paraview 5.11 but the same issue occurred
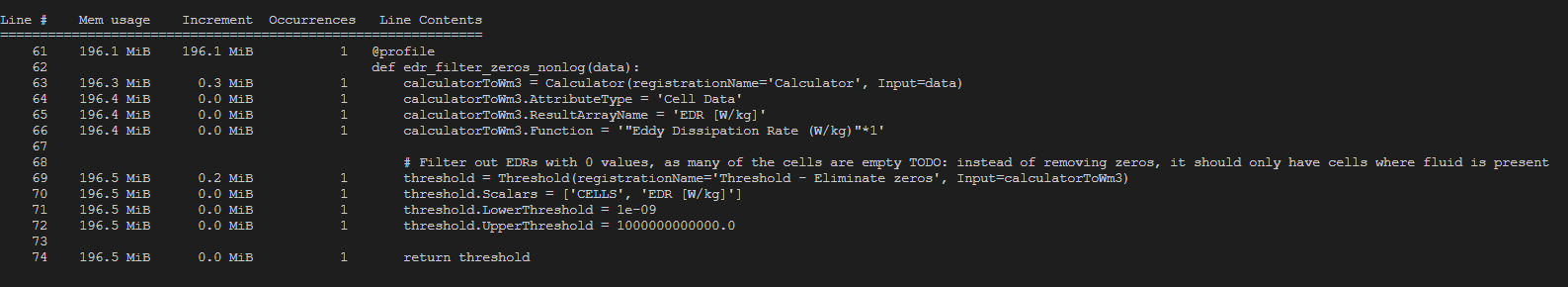
To get some more insights I used a memory_profiler on the relevant functions. The output for loading 1 single .vti file can be seen at the bottom of this post.
Does anyone know how I could fix this issue? If you need any further clarifications on my problem please let me know.
Kind regards,
Ramon
OUTPUT MEMORY_PROFILER PYTHON: