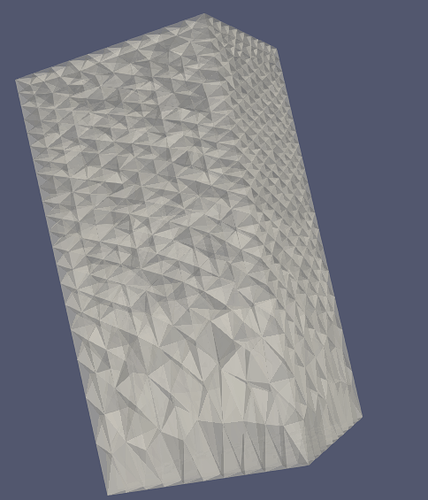
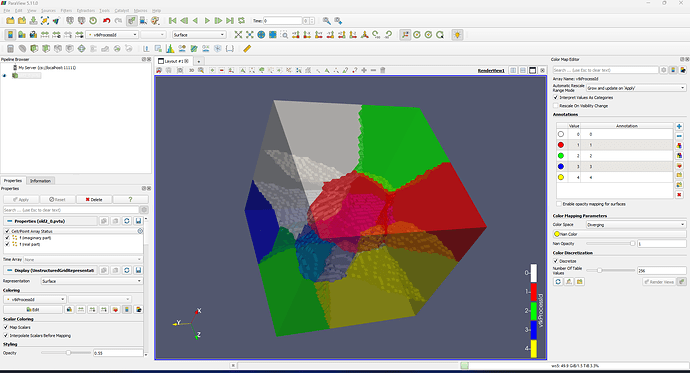
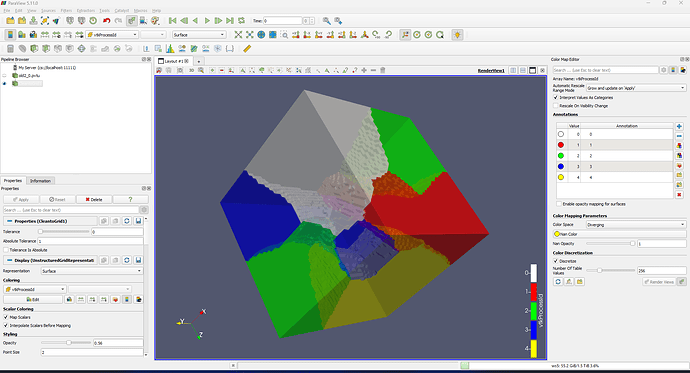
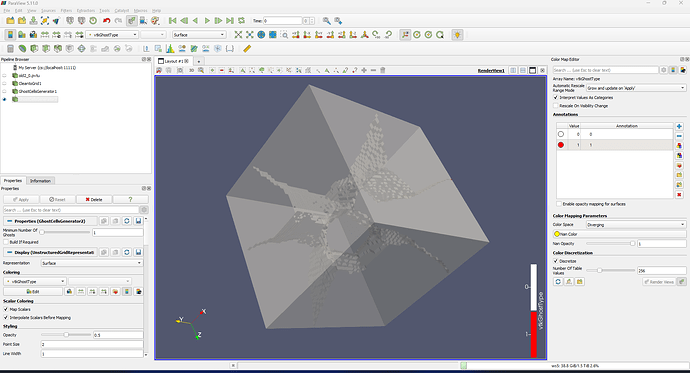
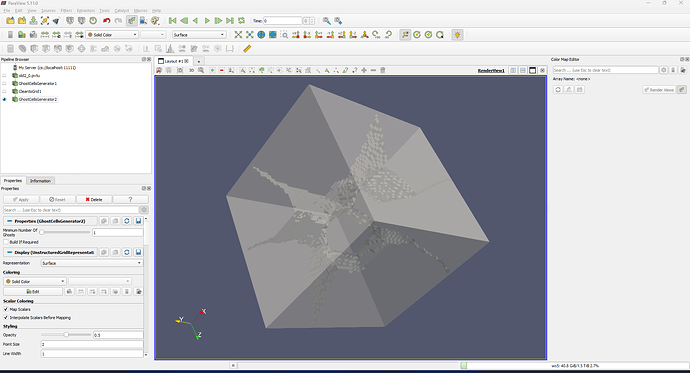
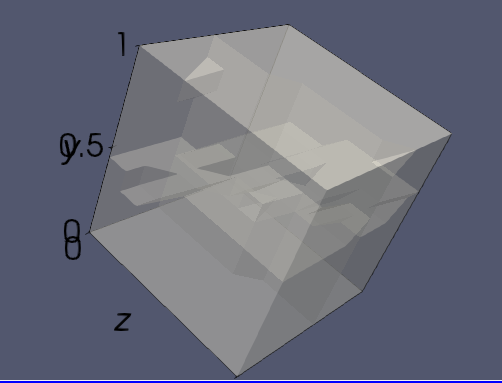
Hi, I am using ParaView to show a box. When I selected the opacity, it showed like below. How to get rid of the tetrahedron face?
It would help if you shared your data, but using the Clean to Grid filter might fix your problem.
Thanks for your suggestion. That works for me.
Hi Kenneth, that works well on a single server process, but when I start the server with mpi, e.g. mpiexec -n 4, the clean to grid won’t merge points on different processes. How to handle this case?
A data set is attached. Thanks!
test.zip (31.3 KB)
You might try the D3 or Redistribute DataSet filters – I feel like one of those fixed this same issue I had several years ago.
@GregVernon The D3 filter fixed this issue. Thank you very much!
@Kenneth_Moreland – while we’re on the topic… what is the reasoning behind the name of the D3 filter? I’d argue that it’s impossible to know what it does from it’s name, it’s as useful as if it were named H8.
Is D3 equivalent to the Redistribute DataSet filter? If so, should D3 be removed / deprecated? At the very least, it seems like it should at least be renamed to Repartition DataSet per its documentation:
<SourceProxy class="vtkDistributedDataFilter"
label="D3"
name="D3">
<Documentation long_help="Repartition a data set into load-balanced spatially convex regions. Create ghost cells if requested."
short_help="Repartition a data set.">The D3 filter is
available when ParaView is run in parallel. It operates on
any type of data set to evenly divide it across the
processors into spatially contiguous regions. The output
of this filter is of type unstructured
grid.</Documentation>
Yang and Greg,
I believe there is a bit of confusion going on here.
The Redistribute Dataset filter does what it sounds like - it redistributes a dataset among multiple back end processors/cores/ranks, when using MPI. You really don’t want to use this to get rid of internal boundaries.
The Clean to Grid filter gets rid of redundant points within the files of one rank. When it is run on a butiltn server, it works on all of the dataset. This should get rid of internal boundaries between files. It will not get rid of boundaries between ranks.
The Ghost Cell Generator filter will effectively remove the boundaries between files on different ranks. Ken created a really good write-up on this in the Self Directed Tutorial (Help/ Self Directed Tutorial). Note that the Ghost Cell Generator filter has had some warts in the past, but is quite clean in master and the upcoming 5.11.0 release.
The D3 filter is older than dirt. It is basically a combination of the Redistribute Dataset filter and the Ghost Cell Generator (well, older versions of same). It is slow and takes lots of memory, but has been a very reliable tool for a lot of years. When the Ghost Cell Generator has had a year or so to prove it’s stability, D3 is slated to fade into the sunset.
@wascott Thank you so much for your detailed explanation!
Hi @wascott, in my recent case, I need to use the filter Ghost Cell Generator twice to remove the boundaries between files on different ranks. Is there some reason behind the scenes?
Hmmm… that is a surprise. You can adjust the number of ghost cell layers, I have never seen a need for this. Anyway, add 2 ghost cells? Alternatively, post an example dataset and include details how to replicate? If the dataset is too big, post screenshots before ghost cell generation, after the first ghost cell generation and again after the second ghost cell generation?
Sorry for the late reply. ![]()
In the new version Paraview 5.11, the problem is still there. I created 5 processes for the server. The screenshot is shown below as well as the data.
test.tar.bz2 (3.6 MB)
Thank you for your time and attention to this matter.